Short-Read Sequencing
BambniTest™
Non-Invasive Prenatal Testing of Single Gene Disorders (NIPT-SGD)
The Future of NIPT
Targeting De Novo Mutations in Fetus
Approximately 1% of all birth defects are caused by single gene disorders¹. Unlike recessive conditions detected via parental carrier screening, a significant portion of these disorders arise spontaneously. 53.1% of patients with single-gene disorders are autosomal dominant conditions, and 87% of these cases resulted from de novo mutations.² The risk of de novo mutations in offspring increases with parental age³, and contributing significantly to congenital heart disease, epilepsy, and intellectual disability⁴.
Many single gene disorders exhibit phenotypic heterogeneity or lack prenatal markers entirely. This creates diagnostic difficulties, as ultrasound may yield inconclusive findings or appear completely normal.
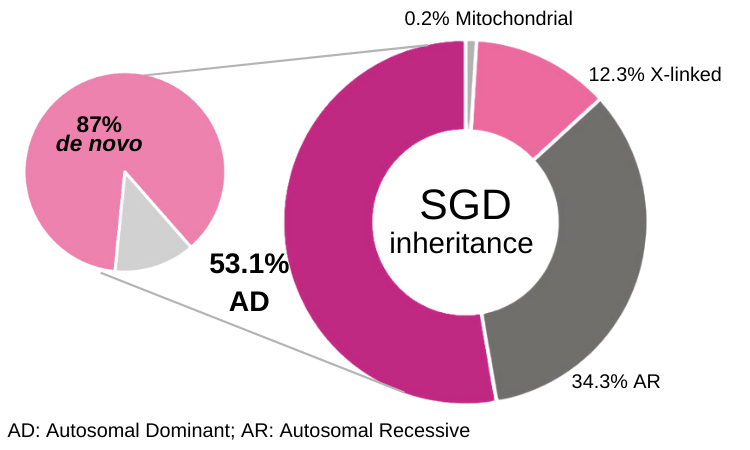
Information derived from reference²
Moving to Target Capture NIPT
BambniTest™ NIPT-SGD represents a shift from genome-wide low-pass NIPT to target capture NIPT. By focusing sequencing power on clinically significant regions (exons, splice sites, and known pathogenic intronic regions for the selected genes), we achieve higher depth and accuracy for small mutations in ccfDNA.
-
Detects de novo abnormalities that cannot be assessed through carrier screening
-
Directly screens for fetal mutations with >99% sensitivity and specificity
-
Provides genetic insight without the risk of miscarriage associated with amniocentesis or CVS
-
Identify severe disorders early, even in the absence of ultrasound anomalies

Selection of Scope
BambniTest™ NIPT-SGD focuses on a defined set of 66 disorders selected for their clinical impact:
-
Incidence rate
-
High penetrance (variants likely to result in a syndrome)
-
Dominant Inheritance, >50% caused by de novo mutations
-
Severe and early on-set (typically appearing <10 years of age)
-
Majority of variants can be screened with NGS technology
-
Conditions at risk of being missed by prenatal ultrasound
Intellectual / Developmental Disabilities
Rett syndrome
CHARGE syndrome
Cornelia de Lange syndrome
Developmental and epileptic encephalopathy
Structural Abnormalities / Others
Osteogenesis imperfecta
Noonan syndrome
Dravet syndrome
Tuberous sclerosis
For details on disease subtypes and genes covered, please refer to the Testing Scope or contact us.
Proven Clinical Utility
In a prospective study⁵, BambniTest™ NIPT-SGD demonstrated robust performance:
-
100% sensitivity & specificity for the test
-
2.06% positive detection rate (72.7% de novo)
-
45.5% positive presented with normal ultrasound parameters
-
Of 445 negative cases with normal delivery and follow-up information, postnatal evaluation (6-15 months) rule out the relevant genetic diseases
537 singleton pregnant women were prospectively enrolled and tested with BambniTest™ NIPT-SGD⁴
End-to-End Technology Transfer
Xcelom Limited and Berry Genomics provide an end-to-end turnkey solution for labs wishing to conduct this test in-house.
-
Laboratory Setup: Full consultation
-
Workflow Integration: SOPs, reagents, and staff training
-
Bioinformatics: Complete software suite for sample management and automated report generation
Platform
Illumina SBS sequencing system
Sample Type
Maternal blood
Operation Time
51/ 55 hours
Automation
Supported
Coverage
Pathogenic (P)/ Likely Pathogenic (LP) SNVs and InDels ≤3bp within 34 genes/ 66 disorders
Note: P/ LP InDels >3bp within 34 genes/ 66 disorders are reported as supplementary
The provided time is based on the Illumina NextSeq 500/NovaSeq 6000 systems and may vary across different labs and systems.
References:
1. Chitty LS. Advances in the prenatal diagnosis of monogenic disorders. Prenat Diagn. 2018;38(1):3-5.
2. Yang Y, Muzny DM, Xia F, et al. Molecular findings among patients referred for clinical whole-exome sequencing. JAMA. 2014;312(18):1870-1879.
3. Jónsson H, Sulem P, Kehr B, et al. Parental influence on human germline de novo mutations in 1,548 trios from Iceland. Nature. 2017;549(7673):519-522.
4. Taylor JL, Debost JPG, Morton SU, et al. Paternal-age-related de novo mutations and risk for five disorders. Nat Commun. 2019;10(1):3043.
5. Zhang H, He J, Teng Y, et al. Non-invasive prenatal testing for dominant single-gene disorders using targeted next-generation sequencing. QJM. 2025;118(5):344-353.