Short-Read Sequencing/ Long-Read Sequencing
Test Management & Analysis Software
The All-in-One Center for Sequencing Analysis and Reporting
Dedicated Software Solutions Powered by a Proven Core
Berry Genomics offers a portfolio of standalone software systems tailored to specific clinical applications—including WESisi™ for WES/Panels, and distinct platforms for NIPT and CNVseq. While each system is deployed separately to suit your lab’s specific focus, they are all powered by the same robust, industry-validated bioinformatics analysis pipeline.
Each dedicated platform features a built-in Laboratory Information Management System (LIMS) designed for that specific workflow. Whether you are running a high-throughput NIPT center or a specialized WES diagnostic unit, our software unifies operations from sample receipt to signed clinical reports in one intuitive interface.
Flexible Deployment
Secure cloud for instant access and ease of use; local server for enhanced data privacy and control
Broad Compatibility
Seamlessly supports external data analysis from major sequencing platforms (Illumina, MGI, etc.) and various commercial capture probes
Full Traceability
Connect patient clinical info, sequencing data, analysis history, and final reports; Every action is logged for auditing
Clinical-Grade Accuracy
Built on industry-standard pipelines enhanced by our patented algorithms, optimized to deliver reliable, reproducible results
Unified Clinical Workflow: From Sample to Report in Single System
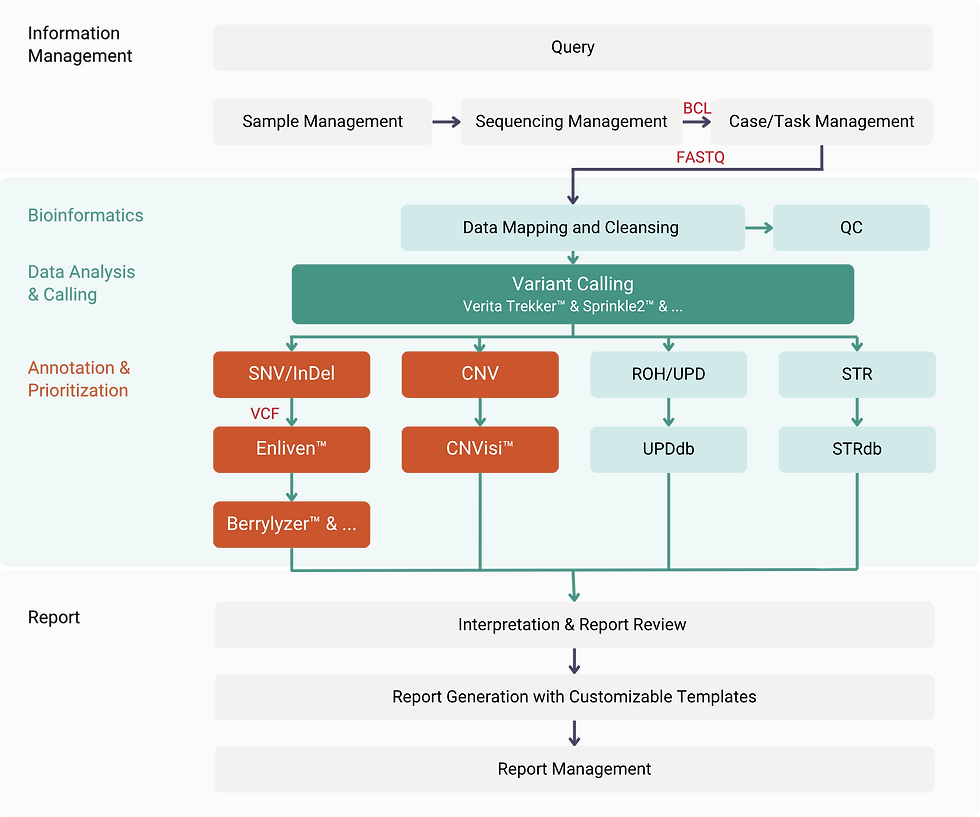
Proprietary Bioinformatics Core
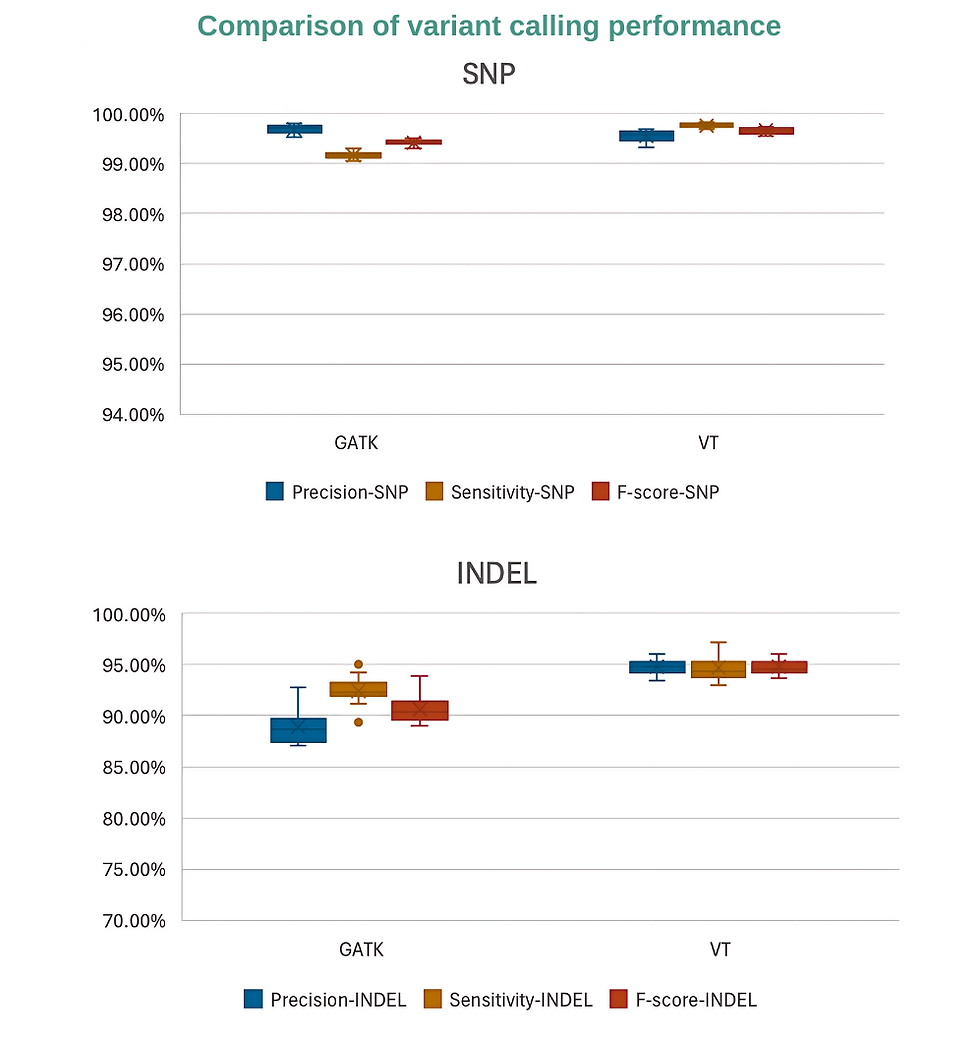
Verita Trekker™: GATK alternative for Small Variant Calling (SNVs/InDels)
Move beyond standard tools. Verita Trekker™ is our proprietary variant detection system designed specifically for small variant calling.
-
Significantly lower memory usage compared to industry-standard GATK without affecting performance
.png)
Resources
Brochure
References:
1. Chen S, Liu C, Luan X, et al. Application and clinical utility assessment of natural language processing-based software for copy-number variants interpretation. J Transl Med. 2025;23(1):1052.